Forward vs Inverse problems: why high performance machine learning usually means little about how the world works
Understanding causality from machine learning is unfortunately usually impossible; life sciences take note
Someone trains a model, the held-out accuracy looks ok, usually “highly significant” but nowhere near product-grade, and the readers/audience is impressed. Then the story quietly shifts from “we can predict” to “we understand the mechanism.” That shift is tempting because the model is doing something real, and “real” starts to feel like “explanatory.” But prediction and explanation are different achievements, and they demand different evidence as I explain here. Throughout this post, ‘works’ will mean two different things. One is ‘predicts new samples like the ones we already see,’ and the other is ‘would change the outcome if we intervened.’
I previously walked through this with neuron interaction estimation; here I want to talk about the general problem which is universal in science.
Forward vs Inverse models
One reason the confusion happens is that we often write the same-looking equation for two different goals.
In the forward (or machine learning) problem, we learn a map from measurements to outcomes, something like
This is a statement about association in the data we observed, not a statement about what would happen if we forced a variable to change. We celebrate when this generalizes to new samples drawn from the same regime.
In the inverse (or causal) problem, we want a story about what generates the data, more like
where Z represents the things that actually drive changes in Y and g() represents how Z actually affects Y, and ε is everything else we did not model. The inverse problem is not ‘harder prediction.’ It is a different target: it asks which parts of the system are levers, meaning if you could set them to a new value, the outcome would move.
Those two equations can both fit the same dataset, yet they answer different questions. The forward question is “can I predict,” while the inverse question is “what would happen if I changed the system.” The problem is that we use the same symbols for both, and then forget which question we are actually answering.
The nature of today’s complex domains
High-dimensional biology (and economics and finance etc) makes the forward problem easier than it looks, in a way that makes the inverse problem harder than it looks. When you measure thousands of variables, much of their variation often rides on a few big latent factors, so the data effectively live near something like
with U low-dimensional and h some complicated embedding. A predictor only needs to learn a function on the observed manifold, and that gets much easier when the effective dimension is small. It does not need to assign credit correctly to each measured feature, as long as it captures the dominant axes well enough to forecast Y. That is why you can get excellent accuracy while still being uncertain about what is doing what (the technical term is collinearity). Held-out test data are usually nearby points on the same surface. Interventions are different: they often move the system in directions the dataset barely explored, so even a perfect predictor on the surface can be unreliable off it.
Why predictive success can coexist with causal ambiguity
Correlations are where people most often slide from forecasting into storytelling. If one variable is causal and another is a tightly correlated proxy, both can predict, and a model will happily use whichever is convenient. When features are highly correlated, many weight vectors predict equally well, so a weight map is often a choice among near-equivalents, not a discovery of the unique driver. A simple way to write the proxy situation is
where η is noise that is small compared to the variation in X1. Now imagine the outcome really follows
In observational data, X2 will look “important” because it shadows X1, even though intervening on X2 might do nothing. The model is not being dishonest; it is answering the forward question you asked it. If you could randomize X2 while holding X1 fixed, prediction would not improve and the outcome would not change, even though X2 looked ‘important’ in observational training.
Here is the linear-algebra version of the same point. Prediction mostly needs Σ, the covariance structure that tells you which directions in the data have real, repeatable variance; causal disambiguation needs Θ=Σ−1, which is an inversion step. Write:
The moment you invert, the small si dominate: directions that were weakly present in the data become massively magnified in the inferred conditional story. That means tiny estimation errors in Σ can produce big changes in causal estimates, even when predictive performance barely budges. In other words, the forward problem can safely ignore the bottom of the spectrum, but the inverse problem is forced to live there (I wrote out some of the math here).
This is also why many popular interpretation tools can feel more decisive than they deserve. They often summarize predictive association, roughly the idea captured by
Here |X1=x denotes what happens if we observe X1 to have the value of x. We read that as if it were a statement about what causes what. The inverse question is different, and it is closer to an intervention statement like the following
Here do(X1=x) denotes what happens if we set X1 to the value of x . The difference is that conditioning compares groups that happen to differ, while intervention describes a world where we force a difference. Those two expectations can be far apart when confounding or tight correlations are present. Observational datasets often never show you the variation needed to separate them, because the system rarely wiggles one variable independently of its entourage. When that independence is missing, the inverse problem is underdetermined even if your predictor is perfect.
Nothing here is anti-prediction or anti-ML. The claim is narrower: predictive success is evidence that information is present, but not evidence that the model has identified manipulable levers. Sometimes this distinction matters in a life-or-death way, like getting a car not to drive into trees. That can support decoding, diagnosis, screening, and control, and it can be scientifically meaningful.
There are real damages
The wreckage from mistaking prediction for causation is everywhere. Pfizer invested heavily (reported around $800 million) in torcetrapib, a CETP inhibitor that dramatically raised HDL cholesterol, because HDL was a strong predictor of cardiovascular outcomes, and then halted the program when the ILLUMINATE trial showed excess deaths and events. The marker moved, but the intervention was not a safe causal lever, and harmful effects unrelated to HDL likely dominated. The homocysteine story followed the same arc: observational data linked it tightly to heart disease, B vitamins reliably lowered it, and then randomized trials showed little to no reduction in major cardiovascular events overall (at most a small stroke signal in some analyses), implying that the observational association did not identify an actionable causal pathway. CARET tested beta-carotene plus retinyl palmitate in high-risk smokers and asbestos-exposed workers because nutrition epidemiology suggested benefit, and lung cancer incidence rose by about 28%. In genetics, GWAS hits frequently tag causal variants through linkage disequilibrium, so risk scores can stratify populations even while the step from association to effector gene and mechanism remains a stubborn bottleneck. In each case, the predictor worked. The causal story did not, or it was never properly identified. Treating correlates as causes contributed to years of wasted effort, large costs, and, in some trials, preventable harm.
Let us do the experiments if we want mechanism
The mistake is letting predictive success quietly impersonate mechanistic understanding. Mechanism begins when you can say what will happen under manipulation, and then show it with the right design. Until then, a strong predictor is best treated as a powerful summary of information, not as a reliable explanation of how the world works. If your evidence is purely predictive, stop making mechanistic claims—or start designing the experiment that would let you.



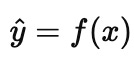
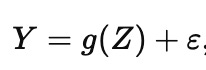
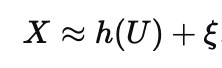
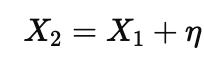
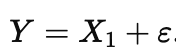
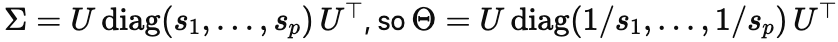
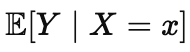
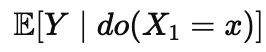
Hi Konrad, I agree with your comments here, but how does this fit with the general practice in NeuroAI, and your claim that there is too much hypothesis testing in NeuroAI and neuroscience? Seems to me that most of NeuroAI is focused on making predictions with naturalistic stimuli, and ignoring experiments that manipulate stimuli to test specific hypotheses about the algorithms and mechanisms that support human and ANN performance. When models are tested on these sorts of experiments (typical in psychology), they generally fail.
Even more problematic, reviewers and editors are so committed to reporting better and better predictions that they are not much interested in publishing studies that falsify ANNs. For example, we can’t get out MindSet Vision toolkit (designed to facilitate this alternative experimental approach to NeuroAI), twice rejected in NeurIPS, once in ICLR, with comments like “It remains unclear what we should do or not do to improve the designs of these models even after benchmarking models on these experiments”.
Or consider the Centaur model published in Nature that catastrophically fails when tested on experiments that manipulate variables (Bowers et al., 2025). For instance, it can often recall 256 digits correctly in a STM digit span task (humans recall about 7 digits), and can output accurate response in a serial reaction time task in 1ms. The editor rejected our commentary, writing:
“…we have come to the editorial decision that publication of this exchange is not justified. This is because we feel that the authors have acknowledged that Centaur is not a theory, nor should it be expected to replicate in extreme situations.”
We then submitted our work to elife, and got the following:
"…as usually with LLM-stuff, and with benchmarks in general, having the criticism hinge on one result that could change with training data/regime is not ideal”.
Yes, some other model might succeed (in two sets of experiments, not one result), but the point is that the model published in Nature does not. And it will be hard to find a better model of human mind if its flaws are difficult to publish and the focus is on predicting held out samples. Falsification is not much appreciated in NeuroAI. Bowers et al. (2023) provide multiple reviews where reviewers and editors and say it is necessary to provide “solutions” to publish; that is, show that ANNs are like humans. Or check out my talk at NeurIPS from a few years ago where I go through many more examples of reviewers and editors rejecting falsification because they are more interested in solutions: https://neurips.cc/virtual/2022/63150
I agree with your post, and think that is undermines many of the claims made in NeuroAI. Am I being unfair to NeuroAI?
Jeff Bowers
Biscione et al. (2025). MindSet: Vision. A toolbox for testing DNNs on key psychological experiments https://openreview.net/forum?id=VkPUQJaoO1&referrer=%5Bthe%20profile%20of%20Jeffrey%20Bowers%5D(%2Fprofile%3Fid%3D~Jeffrey_Bowers1)
Bowers, J. S., Malhotra, G., Adolfi, F., Dujmović, M., Montero, M. L., Biscione, V., ... & Heaton, R. F. (2023c). On the importance of severely testing deep learning models of cognition. Cognitive Systems Research, 82, 101158.
Bowers, J. S., Puebla, G., Thorat, S., Tsetsos, K., & Ludwig, C. J. H. (2025). Centaur: A model without a theory. PsyArxiv https://doi.org/10.31234/osf.io/v9w37_v2
Nice one, thank you.